Albumin Assay – Bromocresol Green method
Practical Documentation:
Aims
To perform serum albumin determinations on samples, explain the principle behind the Bromocresol Green method for albumin measurement and to list the factors that will cause interference with this method.
Principle
Albumin is known for its ability to bind many types of organic compounds, including organic dyes. When albumin selectively binds with Bromcresol Green (BCG) it causes a change in the absorbance maximum of BCG. The intense blue-green complex that is formed has an absorbance max of 670nm. Bromocresol reagent at pH 4.3 is negatively charged. The pI of albumin is 4.7.

For all spectrometric assays, always use a Reagent
blank. It usually contains all diluents and reagent in the reaction
solution, but no sample. Some reagent blanks do contain the sample as well, but
they lack one crucial reagent component needed to produce a colour-yielding
reaction. This is different from the water used to zero a spectrophotometer
(set 100% T).
Materials:
- Albumin Stock Solution (100g/L)
or Cobas calibrator - Serum patient samples (record
the albumin results off track) - Cobas albumin controls
- Bromocresol green reagent (BCG)
– obtain Cobas reagent or make up as follows: - 7.5mg of
BCG - 250mg
EDTA - 250µL
Tween 20 - Bring up
to 25mL with water
Methods:
- Generate a calibration curve of
at least 8 standards (0- 80 g/L) by diluting the Albumin Stock (100 g/L). - In labelled tubes, set up a
calibration, controls and test samples as follows:
| Sample | Water (µL) | Std (µL) | Control (µL) | Sample (µL) | BCG Reagent (µL) |
| Blank | 310 | – | – | – | 300 |
| Calibrators | 300 | 10 | – | – | 300 |
| Control | 300 | – | 10 | – | 300 |
| Serum Sample | 300 | – | – | 10 | 300 |

To set up a standard curve with more points along the usable range, I chose more points in the concentration range 20 – 40 g/L, which in my experience constitutes the bulk of albumin measurements.
3. Mix well read immediately at 630nm and record absorbances
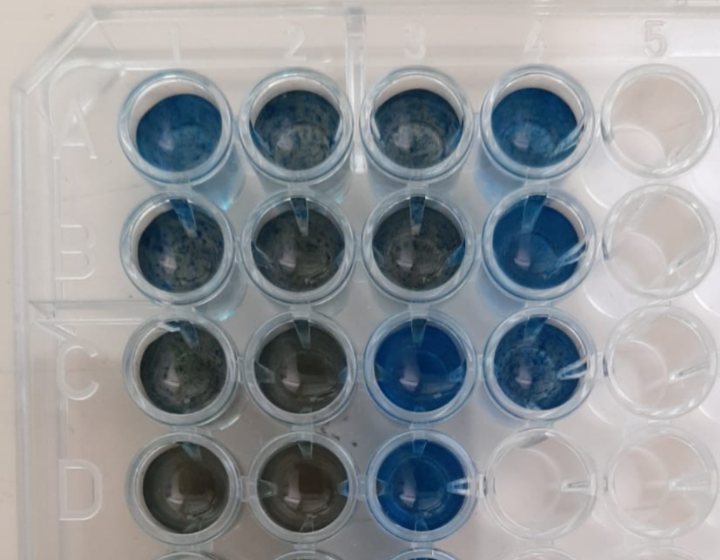
| Layout of wells |
||||
| Dilution as per manual |
Dilution of 1:2 with H2O |
|||
| 1 | 2 | 3 | 4 | |
| A | Standard 0 | Unknown 1 | Standard 0 | Unknown 1 |
| B | Standard 1 | Unknown 2 | Standard 1 | Unknown 2 |
| C | Standard 2 | Unknown 3 | Standard 2 | Unknown 3 |
| D | Standard 3 | Unknown 4 | Standard 3 | Unknown 4 |
| E | Standard 4 | Unknown 5 | Standard 4 | Unknown 5 |
| F | Standard 5 | QC Low | Standard 5 | QC Low |
| G | Standard 6 | QC High | Standard 6 | QC High |
| H | Standard 7 | Standard 7 |


| Absorbance | Absorbance | |
| Standard concentration (g/L) | No dil | Dil 1:2 (150uL reagent mix + 150uL H2O |
| 0 | 0.095 | 0.065 |
| 20 | 0.506 | 0.245 |
| 25 | 0.719 | 0.345 |
| 30 | 0.779 | 0.395 |
| 35 | 0.784 | 0.411 |
| 40 | 0.999 | 0.504 |
| 50 | 1.242 | 0.593 |
| 80 | 1.477 | 0.68 |
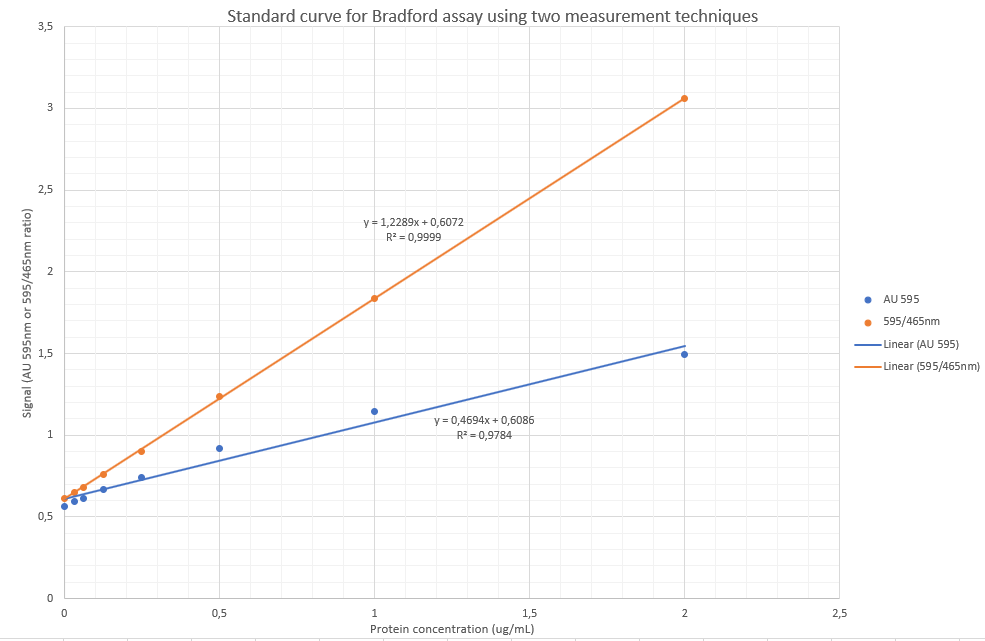
4. Plot a standard curve and determine experimental concentrations of controls and serum samples



These values corresponded to the values when plotting an X-Y plot on Microsoft Excel (see 2 standard curve data plots above).
The figure below illustrates the formula used to determine the unknown concentrations.

Unknown concentrations were calculated as follows:
| Unknowns Sample no. |
Absorbance | Absorbance (Diluted 1:2) |
Calculated concentration | Calculated concentration (Diluted 1:2) |
Reference albumin values (from Roche Cobas 6000) | |
| 1 | 0.893 | 0.48 | Hemolysed | 38.8 | 44.2 | 29.1 |
| 2 | 1.057 | 0.55 | 48.1 | 53.0 | 44.2 | |
| 3 | 0.614 | 0.334 | 23.1 | 26.0 | 22 | |
| 4 | 0.997 | 0.5 | Hemolysed | 44.7 | 46.7 | 38.6 |
| 5 | 0.894 | 0.479 | 38.9 | 44.1 | 33.2 | |
| Lo | 0.843 | 0.453 | 36.0 | 40.9 | 31.9 | |
| Hi | 1.161 | 0.593 | 54.0 | 58.4 | 49.4 |
Compare results to expected results and comment on any differences between the manual BCG vs automated BCP assays.

A possible explanation for this difference is:
1. Pipetting error
2. Interferring substances in the serum which absorb light at 670nm.
3. It may be due to the fact that albumin stock solution was made up with water as opposed to physiological albumin-free serum matrix.
4. Hemolysis – Evidenced by one of the hemolysed samples which clearly measured falsely high.
A haemolysed sample is brought to the laboratory for albumin analysis. Can the sample be used? Discuss.
Yes. Hemoglobin does not absorb light at 670 nm, therefor will not interfere significantly with the analysis. See figure below:

It does however interfere in the following manner:
Hemoglobin decreases the apparent albumin concentration by 1 g/L for each 100 g/L added. Blanking does not correct this interference, and the negative bias is therefore caused by interference with the dye binding rather than hemoglobin color. For the BCP method, a blank correction is required on icteric sera and on grossly hemolyzed and grossly lipemic sera to correct for an underestimation of albumin caused by these agents. Heparin causes a positive interference with BCP and BCG methods. This interference can be eliminated by the addition of hexadimethrine bromide to a concentration of 50 mg/L in the BCP reagent
Kaplan’s Methods
Why is it not desirable to incubate the reaction before measuring the absorbance?
Incubating the sample can give rise to other non-specific binding of analytes in the sample to the chromophore dye and a falsely elevated reading can be obtained.
Some analyzers can measure the absorbance of the BCG reaction within 30 seconds after adding sample. Does this tend to increase the specificity of the reaction?
Yes. The less time there is for other interfering substances to bind to BCG, potentially the more specific it will be to albumin, as albumin is the more specific binding to BCG.
Probably the most promising adaptation of the BCG reaction for albumin analysis utilizes fast reaction readings. Gustafsson reported that measuring the absorbance of the BCG-protein complex at 629 nm at a time shortly after mixing improves the specificity of the assay. Interference by other proteins such as ceruloplasmin and orosomucoid becomes significant at times greater than 5 minutes.
Kaplan’s Methods